About
About
This portal provides access to xQTL results generated from brain and related multi-omic datasets to support search, interpretation, and downstream analysis. It is designed to help researchers explore genetic regulation across genes, variants, and genomic regions in a single, searchable interface.
The portal was developed to make complex xQTL results easier to browse and use, while preserving the transparency and rigor needed for research use. It brings together association results, metadata, and supporting resources so users can move from a query to biological interpretation with less friction.
This resource is maintained by the NIAGADS, FunGenAD-xQTL team in collaboration with contributing investigators and data generators https://adsp-fgc.niagads.org/research. We are grateful to all consortium members, study participants, and analysts whose work made this resource possible.
Team
Acknowledgments
https://dss.niagads.org/datasets/ng00184/
Citation
XXX bioRxiv link
FAQ
1. What data are included in this portal?
This portal provides access to harmonized brain multi-omic xQTL data, including results across multiple omics types, brain regions, and cell types. The current portal highlights 14 brain regions, 7 cell types, 6 omics types, more than 13,000 samples, and over 300 million xQTL associations.
2. How do I search the portal?
You can search by gene, variant, or genomic region using the main search interface. Example queries shown on the portal include genes such as APOE, variants such as rs7412, and region such as chr19:44908000-44909000.
3. What does an xQTL association result mean?
An xQTL association result represents a statistical association between genetic variation and a molecular trait measured in a given dataset or context. In this portal, results are intended to help users explore potential regulatory relationships across genes, variants, and genomic regions.
4. What kinds of xQTLs are available?
Data spans across multiple xQTL categories, including expression quantitative trait loci (eQTLs), histone acetylation quantitative trait loci (haQTLs), methylation quantitative trait loci (mQTLs), protein quantitative trait loci (pQTLs), single-nucleus eQTLs (snuc-eQTLs), and splicing quantitative trait loci (sQTLs). The portal holds the significant associations of these.
5. What studies or datasets are represented in the portal?
The portal describes the resource as integrating brain xQTLs from ROSMAP, MSBB, Knight-ADRC, and MiGA. These datasets are processed in a harmonized format to support cross-dataset exploration.
6. Is there a genome browser or data explorer?
Yes. Use the Browse (a.k.a. ‘View in Genome Browser’ tab) or track view to explore data summaries, genomic contexts, and xQTL tracks interactively without a specific gene/variant query
7. How do I download data from the portal? 8. What file formats are available for download? 9. How should I cite this portal and its data? 10. Who do I contact if I have questions or find a problem? Contact: help@niagads.org
Updates:
V1.0 – first version of NIAGADS xQTL data is released on 03/30/2026.
The Alzheimer's Disease Sequencing Project (ADSP) Functional Genomics Consortium (FunGen-AD) website FunGen-xQTL Computational Protocol website FunGen-xQTL Computational Protocol [Github] xQTL protocol core modules [pipeline source code] hipFG: high-throughput integration and processing for functional genomics data
Choose from three approaches based on your needs:
We have two kinds of files
a) The metadata is available in both JSON and TSV formats.
b) All xQTL files are available in xxx format and these column names:
Chr: Chromosome
Start: Start of the position
Users should cite the DOI xxx , as well as put the contents from the NIAGADS acknowledgment page into the acknowledgment session when using these data (xxx).
Please contact help@niagads.org for questions.
ADSP Functional Genomics Consortium (FunGen-AD)
FunGen-AD Resources
FunGen-xQTL Computational Protocol
Code repositories
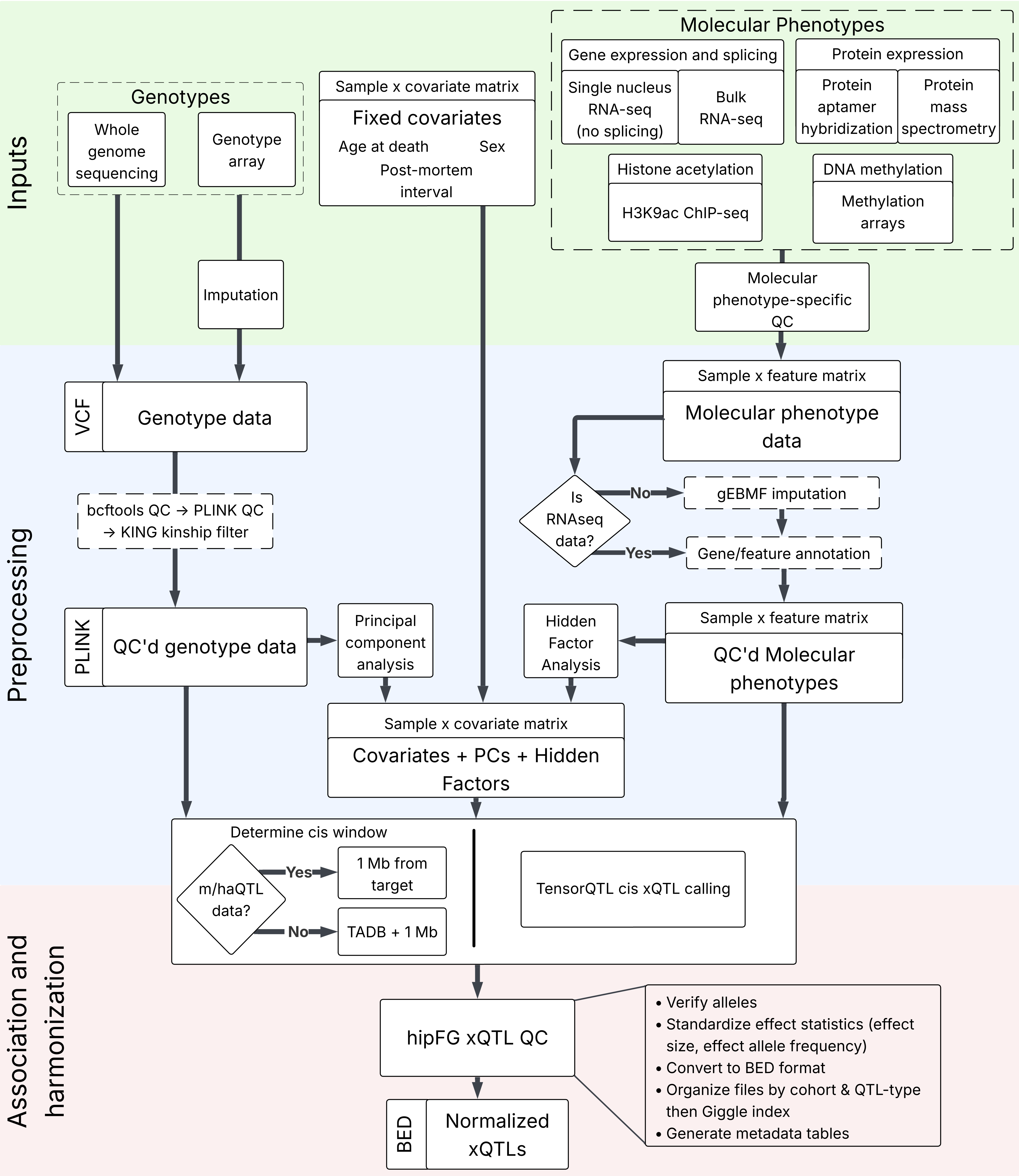
FunGen-AD xQTL data flow